Reactions: CRISPR systems with new properties "resurrected" from extinct bacteria
An international study led by Spanish researchers has succeeded in reconstructing the ancestors of the CRISPR-Cas system present in extinct bacteria from up to 2.6 billion years ago. The reconstructed systems work and are more flexible than the current ones. According to the authors, this could open up new avenues for gene editing. The results are published in the journal Nature Microbiology.
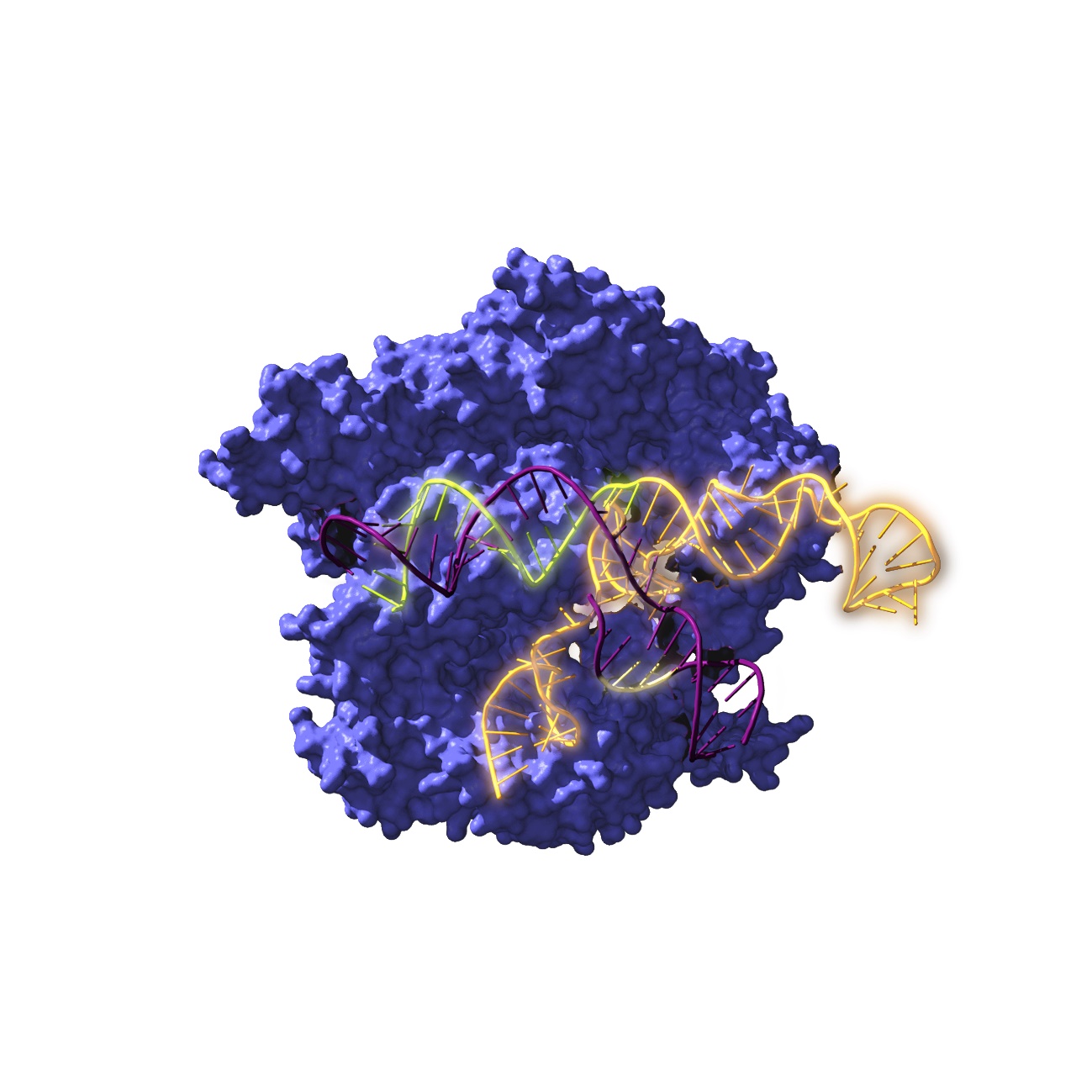
Image of Cas9, an endonuclease enzyme associated with the CRISPR system, acting on the target DNA / Antonio Reifs (CIC nanoGUNE).
Moreno-Mateos - Crispr (EN)
Miguel Ángel Moreno-Mateos
Tenured scientist CSIC & PI, Andalusian Center for Developmental Biology, CSIC-Pablo de Olavide University
It is a genuinely original work with an imaginative approach to ultimately understand the evolution of the CRISPR-Cas9 system over billions of years.
Improving CRISPR-Cas technology requires a good understanding of how these systems work from a biochemical and structural point of view. Knowing how they have changed over the course of evolution will allow us to approach these improvements from a new point of view, with potential applications in biotechnology and biomedicine.
The study lays a foundation on which we can now continue working and be able to use this knowledge in tangible biotechnological applications. In any case, these basic studies are fundamental for the advancement of more applied sciences.
Toro - Crispr
Nicolás Toro
Nicolás Toro, CSIC Research Professor.
Bacteria harbour a large number of systems to defend themselves against foreign biological attacks, including viruses, which have evolved over millions of years and are of great interest today because of their potential usefulness in correcting genetic defects and diseases affecting higher organisms, including humans. Among these systems, perhaps the best known is CRISPR-Cas, an adaptive immunity system that allows bacteria to collect and store the history of successive attacks and thus lay the foundations for protection and defence against those of the same nature that may occur in the future.
The study recently published in the prestigious journal Nature Microbiology, led by Spanish scientists, succeeds in reconstructing the evolutionary history of one of the most famous of these systems, the one associated with the Cas9 effector protein. The study of the now extinct ancestors of this current protein reveals an evolution from systems that were originally more versatile, both in the recognition of target sequences and in their nature, probably single-stranded DNA and RNA.
This study raises new questions about what the initial function of these primitive CRISPR-Cas systems was, and their versatility suggests that these reconstructed proteins from a bygone era could be very useful in genome editing today, opening up a new future for the use of this technology. However, the use in general and in particular of these new genetic technologies in humans undoubtedly raises many concerns from an ethical and social point of view, and studies are needed before they can be used effectively, especially the evaluation of unwanted activity elsewhere in the genome.
Technology based on the primitive CRISPR-Cas systems identified in this study, as well as the exploration of the diversity of other bacterial defence systems, may represent a revolution in the advancement of medicine for humankind.
Alonso-Lerma et al.
- Research article
- Peer reviewed
- Experimental study
- In vitro