Reactions to preliminary results of a new universal flu vaccine prototype
A study published in the journal Science reports preclinical results of a new vaccine model that is intended to work against all types of influenza. The prototype, which uses mRNA-based technology, includes antigens from all 20 known influenza subtypes.

Adolfo García Sastre - gripe EN
Adolfo García-Sastre
Director of the Institute for Global Health and Emerging Pathogens at Mount Sinai Hospital in New York
This is a very interesting study. It demonstrates the ability to be able to develop multivalent mRNA vaccines that are able to immunise against 20 or perhaps more different antigens at the same time. In this case, these are influenza virus antigens that encompass all possible influenza virus subtypes and variants, including those with pandemic potential.
Current influenza vaccines do not protect against influenza viruses with pandemic potential. This vaccine, if it works well in people, would achieve this.
The studies are preclinical, in experimental models. They are very promising and, although they suggest a protective capacity against all subtypes of influenza viruses, we cannot be sure until clinical trials in volunteers are done.
"I am a patent inventor of influenza virus vaccines in clinical development, which use a different strategy than the one used in this study to obtain protection against all influenza viruses".
Nistal - Gripe (EN)
Estanislao Nistal
Professor of Microbiology at the Faculty of Pharmacy
The study is highly relevant and well conducted, providing very interesting data on the possibility of generating a universal vaccine against the different influenza viruses.
Influenza disease in humans is mainly caused by influenza A and B viruses. Currently, the influenza A virus in humans can be further subclassified into H1N1 and H3N2 subtypes, depending on the class of the haemagglutinin (there are 18 different versions in total) and neuraminidase (there are 11 different versions) proteins that the viral particles have on their surface. Also influenza B viruses can have two different versions of haemagglutinin on their viral particles. There are numerous subtypes of influenza A viruses infecting animal species other than humans that contain the different versions of haemagglutinin and neuraminidase mentioned above.
It is enormously complex to formulate a vaccine against influenza viruses that can protect us, not only against currently circulating viruses in humans, but also against potential viruses that could produce a future pandemic. Such a vaccine is the so-called universal influenza vaccine.
The researchers present a strategy similar to that used to generate the messenger RNA vaccine against SARS-CoV-2, but in which they introduce messenger RNA from the 20 versions of the haemagglutinins of type A and B influenza viruses that could give rise to a virus with the potential to infect us. The results show that this vaccine is able to induce a robust antibody-mediated response in mice and ferrets (animal models widely used to study influenza) against different subtypes of influenza viruses, including viruses that are significantly different from the sequences included in the vaccine.
All of this implies the potential for an easily and rapidly constructed universal vaccine that could be of great use in the event of a pandemic outbreak of a novel influenza virus. Although not discussed in the article, this vaccine could also be of great use in preventing influenza in animals that may suffer from it, and reducing the risk of zoonosis among animals in a global health context.
The article does not yet present data on the possible advancement of this vaccine to a next phase in humans, where not only efficacy, but also adverse effects, dosage or short- and long-term immunity should be demonstrated.
Another limitation is that they need to further study the role of T-lymphocytes in disease protection. Activation of CD4 T cells is important for an optimal humoral antibody response.
Raúl Ortiz - vacuna universal gripe EN
Raúl Ortiz de Lejarazu y Leonardo
Professor of Microbiology, scientific advisor and director emeritus of the National Influenza Centre in Valladolid
The search for a universal influenza vaccine began in 2012-13, although there had been different approaches before that. Among the latest is trying to discover and obtain antigens (epitopes) that are present in the majority of variants of an influenza subtype (conserved epitopes). In this way a universal response can be achieved and even if the variable epitopes change, the response to the conserved epitopes will remain.
The new study is well conceived and comprehensively conducted. Science is a journal whose publications are of the highest standard.
The most important novelty is that it uses many antigens of different haemagglutinin subtypes (all existing ones including those from bats) instead of going to conserved regions of one or a few antigens. This used to be more difficult than it is now. Current mRNA vaccine platforms allow the inclusion of many mRNAs that will induce many different proteins giving a multivalency and breadth of response that was not easy to achieve with protein platforms in the past.
The main limitation is that it is done in mice and ferrets, very good animal models for flu, but animal models. So with sarcasm (always healthy in science), mice and ferrets around the world should congratulate themselves because they now have a universal flu vaccine for themselves.
In all seriousness, there is a long, sometimes insurmountable, way to go from the animal model to humans. The type of response, the extent of the response, the persistence, etc. are not similar.
The first [phase 1] human trial of a universal influenza vaccine was published two years ago, but people were enraptured by the new coronavirus.
Víctor Jiménez - vacuna universal gripe EN
Víctor Jiménez Cid
Professor of the Department of Microbiology and Parasitology, Faculty of Pharmacy, Complutense University of Madrid
Science today publishes a paper presenting preclinical trials in mice and ferrets of an 'icosavalent' mRNA vaccine against influenza. This is the first high-impact publication to present a successful strategy for a "universal" mRNA-based vaccine against influenza. The formulation includes modified RNAs formulated into lipid nanoparticles, the same technology used by Moderna in the development of the widely distributed SARS-CoV-2 vaccines.
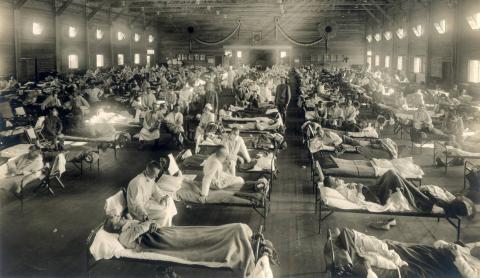
The vaccine includes the 18 known haemagglutinin spicule types of influenza A viruses (H1-H18) plus two for influenza B viruses. For reference, the tri- or tetravalent influenza vaccine we are currently using contains one A/H1, one A/H3 and one or two B viruses. The seasonal influenza A viruses circulating in the human population are only H1N1 and H3N2. Why include other H antigenic types in the vaccine? Firstly, A viruses are zoonotic, and although the other types do not circulate in the human population, they do circulate in other animals, such as H5 H7 and H9 in birds. This implies that new pandemic viruses can emerge if one of these types engages in "antigenic jumping" by generating a new A virus that combines the genes of animal viruses with those of viruses circulating in humans. This is what happened in 2009, 1968 (Hong Kong flu), 1957 (Asian flu) and in the terrible 1918 pandemic that killed at least 50,000,000 people.
This type of vaccine would therefore prevent, in addition to seasonal influenza, the spread of avian influenza in humans, which has a mortality rate of around 30%, and possible new emerging pandemic viruses. Most importantly, the authors of the study show that their cocktail generates antibodies to low-variable regions of the HA spicule, which are found in the structure of the 'stem'. The enormous mutational capacity of influenza viruses means that the virus "changes its face" from one year to the next. In other words, even if we have built up immunity to the previous winter's flu, this winter our antibodies will no longer recognise the new virus. This "antigenic drift" is similar to what happens with the sublineages of the SARS-CoV-2 omicron variant: as soon as the virus changes antigens, our immune memory no longer recognises it and we can be reinfected with the new variant.
We have known for a long time that the influenza virus does this much faster than the coronavirus and this forces us to reformulate the vaccine every year according to the data that an international epidemiological surveillance service handles to predict what will be the presumably most effective vaccine composition for the winter season. But all that chameleon-like ability of the virus is in the "head" of the HA spicule of the virus. If we can neutralise the invariant region of the stem, we would have a "universal" vaccine, a weapon against the virus's ability to vary. The authors of this study found in their preclinical trials that using this strategy, experimental animals develop neutralising antibodies to the stem, as well as a wide range of antibodies to the 20 different "heads" of the virus' haemagglutinin.
In short, the strategy shows good protection in experimental animals against infection, generating antibodies against all types, something that would be very difficult to achieve with conventional vaccines, and protecting the animals against infection by several types of H1N1. It appears that the presentation of the antigen to our immune system is much more effective in mRNA-based formulations, which force our cells to produce the antigen in situ, than in classical formulations based on direct inoculation of the antigen. Thanks to the technological development forced by the emergency situation created by covid-19, the formulation of vaccines and other mRNA-based drugs is entering a golden age that may lead to a revolution in the prevention and treatment of infectious diseases and other pathologies.
- Research article
- Peer reviewed
- Experimental study
- Animals